Two years ago, the Gordon and Betty Moore Foundation’s Marine Microbiology Initiative pursued an ambitious opportunity to accelerate the study of microbes in the ocean using genetic tools. Marine microbes (comprised of bacteria, archaea and protists) and the viruses that infect them account for 98 percent of the ocean’s biomass and are responsible for driving the carbon cycle of the planet, but scientists are still trying to understand how these abundant organisms function. New tools and technologies have the potential to help transition the field of microbial oceanography from an observational to experimental science, giving researchers the ability to probe these organisms more deeply and ask fundamental questions about their roles in the ocean. How do they live and interact with one another? How do they drive nutrient cycles? Genetics is a predominant cornerstone of the biological sciences and expanding its use in marine microbial ecology promotes an important path for addressing these questions.
The case for model systems and focusing on protists
In 2015, we supported more than 100 scientists from around the world to develop genetic tools to understand what makes marine microbes tick. These researchers were challenged to identify marine protists – single-celled microbes which contain a nucleus – that are amenable to genetic modification and can serve as models for further study.
Protists are among the most diverse group of microbes on earth. They are found in nearly every ecosystem, but little is known about how they succeed in so many environments. Inspired by examples from existing robust model systems, such as the mammalian gut bacterium E. coli, the cohort of scientists are creating genetic tools to reveal how protist genes function. However, there are key challenges they must tackle first, including being able to culture these enigmatic protists in the lab and to manipulate them at the molecular level.
“The most useful model organisms are genetically tractable and, increasingly, amenable to genome engineering techniques. However, gene delivery and genome engineering techniques are not easily transferred across the tree of life; they must be tailored to the biology of each species,” said foundation grantee Dr. Nicole King, professor of genetics, genomics and development at University of California, Berkeley. “In many cases, the most interesting organisms have unusual biology and are studied by relatively small numbers of scientists, meaning that there can be a lot of inertia working against developing them into new models.”
Dr. King and her team study choanoflagellates, protists that are the closest living single-celled relatives of animals. King and researchers in her lab have had to find new strategies for delivering DNA into the cells of these protists and identifying genetic on/off switches, called “promoters,” that work to drive gene expression. In some cases, the team uses promoters to turn on genes encoding fluorescent proteins to give them a visual way of confirming they can properly introduce foreign DNA into choanoflagellate cells.
“Failure at any step of the process would have meant that we had no positive read-out and therefore wouldn’t know where to start troubleshooting. Fortunately, it all came together, but we couldn’t have pursued such a high-risk project without the support of the Moore Foundation,” added King.
Signs of progress and additional investment to continue research
Despite the complexities associated with developing model systems for marine microbes, progress is being made and continued investment in this field of research is still needed. Through our initiative, scientists were provided with an additional $4.5 million to continue development of experimental model systems for protists, bringing the total to $12.5 million. This effort is a core part of the initiative’s focus to advance understanding of marine microbial communities by enabling researchers to uncover the scientific principles that govern the interactions among microbes and microbially mediated nutrient flow in the sea.
“There is not a lot of funding for developing genetic tools for marine microbes, yet even a modest investment yields results that can revolutionize the field,” stated Adam Jones, Ph.D., program officer for the foundation’s Marine Microbiology Initiative. “We want to give scientists support to create new tools that can answer the unknown – how protists interact with their environment and their microbial neighbors – thereby generating new ecological and evolutionary hypotheses for ocean investigation. We are excited by results to date and that is why we are continuing to bet on scientists to develop tools and methods that have never existed for this field.”
In addition to King, others are making progress in the experimental model systems effort. Dr. Elena Casacuberta, a foundation grantee and researcher at the Institut de Biología Evolutiva in Barcelona, Spain is working on tools for two protists: Abeoforma  whisleri and Corallochytrium limacisporum. These protists are also closely related to animals and serve as valuable systems for understanding how animals evolved from a common single-celled ancestor. The tools she and her colleagues are developing will not only be valuable for other scientists, but will also unveil cellular biology and functional characteristics relevant to understanding the lifestyles of these protists, their evolutionary history and their importance in the marine environment.
whisleri and Corallochytrium limacisporum. These protists are also closely related to animals and serve as valuable systems for understanding how animals evolved from a common single-celled ancestor. The tools she and her colleagues are developing will not only be valuable for other scientists, but will also unveil cellular biology and functional characteristics relevant to understanding the lifestyles of these protists, their evolutionary history and their importance in the marine environment.
“Our developments will offer new protocols for the community to apply directly or as an initial step to try in other organisms,” said Casacuberta. “Moreover, while developing the tools we are unveiling special characteristics or behaviors from several protists that might be relevant for other marine organisms in the community.”
Dr. Julius Lukeš, another foundation grantee and a professor at the Czech Academy of Sciences, has made substantial progress working with the marine protist Diplonema papillatum. Recent environmental surveys have revealed Diplonema to be one of the 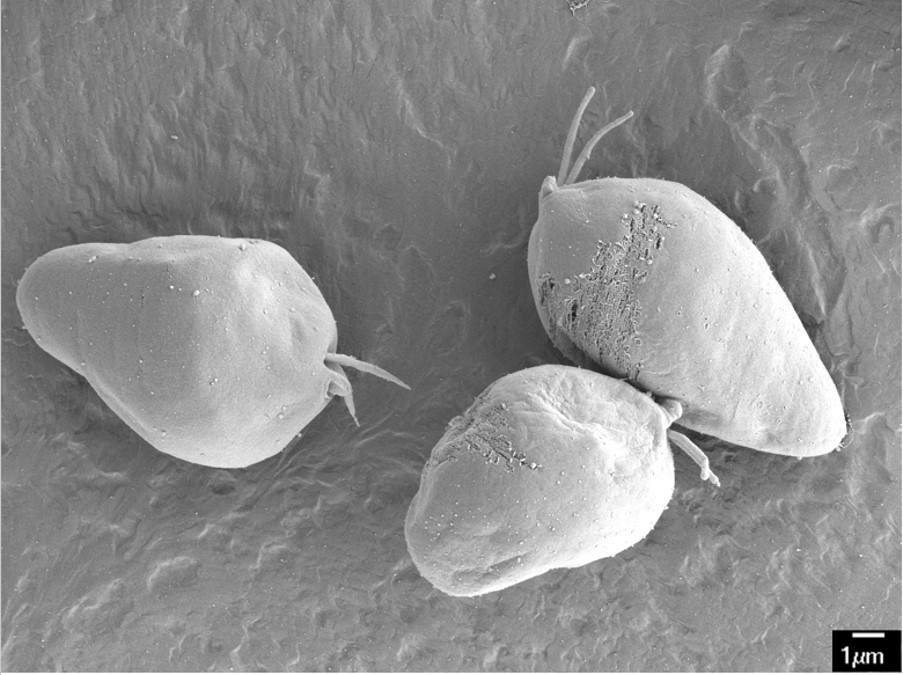 most abundant and diverse groups of protists in the ocean while remaining among the least understood. It is believed that many diplonemids are parasites of other organisms. Lukeš and his collaborators, Drs. Gertraud Burger at the University of Montreal and Patrick Keeling at the University of British Columbia, have been successful in getting artificially constructed gene segments into Diplonema cells.
most abundant and diverse groups of protists in the ocean while remaining among the least understood. It is believed that many diplonemids are parasites of other organisms. Lukeš and his collaborators, Drs. Gertraud Burger at the University of Montreal and Patrick Keeling at the University of British Columbia, have been successful in getting artificially constructed gene segments into Diplonema cells.
“Diplonemids have been shown to be extremely diverse and very abundant in the world ocean but overlooked until now,” stated Lukeš. “No molecular and cell biology has been done with them because there haven’t been advanced methods established for Diplonema. If we and others can demonstrate that diplonemids are genetically manageable, it will not only accelerate the field but open an entirely new field.” Lukeš added that unless methods exist for scientists to knock genes in and out, tag them and more, Diplomena cannot be studied as deeply and will remain marginalized.
Community building among model systems researchers
If we are to accelerate this field, it is important to have open access to the experimental model systems scientists are developing. To help make this possible, we are supporting and partnering with the online, open-access platform protocols.io. Through this repository, scientists can share laboratory protocols, lessons they have learned, and explore potential collaborations.
“Enabling scientists to share protocols and exchange ideas centered on protist genetics freely will help advance scientific research and reach non-protist geneticists who may be working on similar challenges in different systems, such as yeast and mice,” said Sara Bender, Ph.D., program officer for the foundation’s Marine Microbiology Initiative. “There are ripe opportunities for model systems communities, in the ocean and elsewhere, to share lessons learned and brainstorm on best practices for tackling tool development in these ubiquitous yet unfamiliar organisms.”
Our vision is to equip the research community with powerful and essential tools so that scientists can determine how marine microbes interact within their environment and with one another. Through this new knowledge, we have the potential to better understand the role of marine microbes in driving how our ocean’s ecosystems function.

Message sent
Thank you for sharing.